LIG_Dynein_DLC8_1
| Accession: | |
|---|---|
| Functional site class: | Dynein (Light Chain) binding motifs |
| Functional site description: | Motifs are found in the intermediate chain of cytoplasmic Dynein complex (DIC) and many other proteins that bind to light chains. The best characterised motif is the KxTQT motif that binds to DLC-8 (highly conserved 8-kDa light chain of Dynein). |
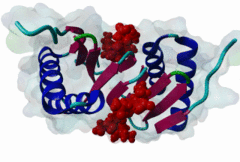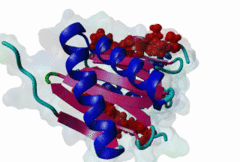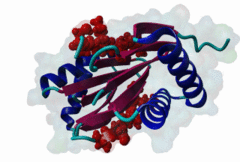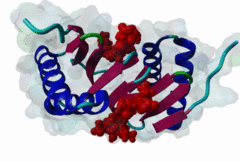
| ELM Description: | [KR]xTQT is a conserved motif found in the intermediate chain of Dynein complex (DIC) and many other proteins. It participates in the precise assembly essential to dynein motor proper functioning. It binds to DLC-8 (highly conserved 8-kDa light chain of Dynein) by beta-augmentation as an extended beta-strand. The motif is also found in viral proteins, suggesting that DLC-8 might be involved in dynein mediated retrograde transport of these proteins along the microtubules in infected cells. |
| Pattern: | [^P].[KR].TQT |
| Pattern Probability: | 0.0000226 |
| Present in taxons: | Arabidopsis thaliana Drosophila melanogaster Drosophila pseudoobscura Eukaryota Homo sapiens Mus musculus Plasmodium falciparum Saccharomyces cerevisiae Schizosaccharomyces pombe Takifugu rubripes |
| Interaction Domain: |
Dynein_light (PF01221)
Dynein light chain type 1
(Stochiometry: 1 : 1)
|
Dynein is a large, multisubunit molecular motor that translocates cargoes toward the minus end of microtubules. It contains 2 heavy chains (DHC), several intermediate chains (DIC) and light intermediate chains (DLIC) and a number of light chains (DLC). Specific assembly of the various subunits has to be precise to ensure functioning of Dynein. The intermediate chain (DIC) of cytoplasmic dynein binds to the highly conserved 8-kDa light chain (DLC-8) through a highly conserved 10 aa stretch (reads SYSKETQTPL). A yeast 2-hybrid screen using DLC-8 as bait showed that a wide variety of additional partner candidates exist and that DLC-8 binding proteins contain the consensus sequence (K/R)XTQT. The motif interacts with the common target-accepting grooves of the DLC-8 dimer. The motif binds by beta-augmentation to form a beta-stranded structure in the DLC-8-target peptide complex. The backbone hydrogen bonds are to one DLC-8 monomer while the Q sidechain makes polar interactions with the second molecule. Because it binds to a large number of functionally unrelated proteins, DCL-8 is believed to act as a multifunctional regulatory protein. A variant motif in nNOS - GIQVD - is found to bind in a similar manner to DLC-8. In yeast, a repeating (VLT)QT motif in the nuclear pore protein nup159 binds to the DLC Dyn2. A KxTQV motif in DIC binds to the DLC TcTex1. It therefore seems likely that other variants of the Q-based DLC interaction motifs will be found. |
-
The 8-kDa dynein light chain binds to its targets via a conserved
(K/R)XTQT motif.
Lo KW, Naisbitt S, Fan JS, Sheng M, Zhang M
J Biol Chem 2001 Apr 27; 276 (17), 14059-66
PMID: 11148209
-
Structural basis of diverse sequence-dependent target recognition by the 8
kDa dynein light chain.
Fan J, Zhang Q, Tochio H, Li M, Zhang M
J Mol Biol 2001 Feb 9; 306 (1), 97-108
PMID: 11178896
-
Identification of novel cellular proteins that bind to the LC8 dynein
light chain using a pepscan technique.
Rodriguez-Crespo I, Yelamos B, Roncal F, Albar JP, Ortiz de Montellano PR, Gavilanes F
FEBS Lett 2001 Aug 17; 503 (2), 135-41
PMID: 11513870
-
Molecular basis for the functional interaction of dynein light chain with
the nuclear-pore complex.
Stelter P, Kunze R, Flemming D, Hopfner D, Diepholz M, Philippsen P, Bottcher B, Hurt E
Nat Cell Biol 2007 Jul; 9 (7), 788-96
PMID: 17546040
-
Structural and thermodynamic characterization of a cytoplasmic dynein
light chain-intermediate chain complex.
Williams JC, Roulhac PL, Roy AG, Vallee RB, Fitzgerald MC, Hendrickson WA
Proc Natl Acad Sci U S A 2007 Jun 12; 104 (24), 10028-33
PMID: 17551010
-
Structure and dynamics of LC8 complexes with KXTQT-motif peptides: swallow
and dynein intermediate chain compete for a common site.
Benison G, Karplus PA, Barbar E
J Mol Biol 2007 Aug 10; 371 (2), 457-68
PMID: 17570393
(click table headers for sorting; Notes column: =Number of Switches, =Number of Interactions)
| Acc., Gene-, Name | Start | End | Subsequence | Logic | #Ev. | Organism | Notes |
|---|---|---|---|---|---|---|---|
| Q15326 ZMYND11 ZMY11_HUMAN |
451 | 457 | ASSPRMLHRSTQTTNDGVCQ | TP | 5 | Homo sapiens (Human) | |
| O43521 BCL2L11 B2L11_HUMAN |
110 | 116 | SPAPMSCDKSTQTPSPPCQA | TP | 4 | Homo sapiens (Human) | |
| O43521-2 BCL2L11 B2L11_HUMAN |
50 | 56 | SPAPMSCDKSTQTPSPPCQA | TP | 5 | Homo sapiens (Human) | |
| P22363 P PHOSP_RABVC |
142 | 148 | PPRRSSEDKSTQTTGRELKK | TP | 1 | Rabies virus CVS-11 | |
| O88485 Dync1i1 DC1I1_MOUSE |
149 | 155 | PREVVSYSKETQTPLATHQS | TP | 6 | Mus musculus (House mouse) | |
| Q13409 DYNC1I2 DC1I2_HUMAN |
156 | 162 | PREIVTYTKETQTPVMAQPK | TP | 1 | Homo sapiens (Human) | |
| Q24246 sw DYIN_DROME |
128 | 134 | PKETLVYTKQTQTTSTGGGN | TP | 2 | Drosophila melanogaster (Fruit fly) | |
| P40688 swa SWA_DROME |
289 | 295 | HIRSATSAKATQTDFLVDTI | TP | 8 | Drosophila melanogaster (Fruit fly) | |
| Q9U9I5 swa SWA_DROPS |
281 | 287 | FCPPASVPKATQTDHELLSH | TP | 0 | Drosophila pseudoobscura pseudoobscura |
Please cite:
ELM-the Eukaryotic Linear Motif resource-2024 update.
(PMID:37962385)
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement