LIG_MAD2
| Accession: | |
|---|---|
| Functional site class: | MAD2 binding motif |
| Functional site description: | Mitotic spindle checkpoint protein MAD2 binding motif |
| ELM Description: | Mitotic spindle checkpoint protein MAD2 binding motif |
| Pattern: | [KR][IV][LV].....P |
| Pattern Probability: | 0.0001011 |
| Present in taxons: | Drosophila melanogaster Eukaryota Homo sapiens Mus musculus Rattus norvegicus Saccharomyces cerevisiae Xenopus laevis |
| Interaction Domain: |
HORMA (PF02301)
HORMA domain
(Stochiometry: 1 : 1)
|
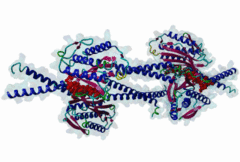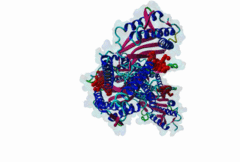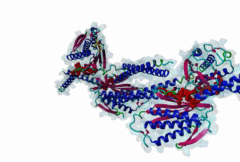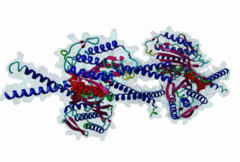
The spindle assembly checkpoint monitors sister chromatid bi-orientation in mitosis and prevents loss of sister chromatid cohesion in the presence of a demaged spindle. Mad2 and other proteins, such as Mad1 and Bub1, associate with kinetochores in prophase and prometaphase to monitor attachment of sister chromatids to the spindle. The target of the spindle checkpoint is the anaphase-promoting complex (APC), a multi-subunit ubiquitin ligase, whose activity determines loss of sister chromatid cohesion and metaphase-anaphase transition. APC inhibition requires a direct interaction of Mad2 with Cdc20, a positive regulator of the APC. Mad2 also interacts with Mad1. Cdc20 and Mad1 complexes with Mad2 are exclusive. The Mad2 binding sites on Mad1 and Cdc20 have been mapped using genetic and biochemical approaches. It has been shown that these unrelated proteins share an 10-residues Mad2 binding motif. This suggests that Mad2 is transferred from one ligand to the other via competition. |
-
The Mad2 spindle checkpoint protein undergoes similar major conformational
changes upon binding to either Mad1 or Cdc20.
Luo X, Tang Z, Rizo J, Yu H
Mol Cell 2002 Jan; 9 (1), 59-71
PMID: 11804586
-
Crystal structure of the tetrameric Mad1-Mad2 core complex: implications
of a 'safety belt' binding mechanism for the spindle checkpoint.
Sironi L, Mapelli M, Knapp S, De Antoni A, Jeang KT, Musacchio A
EMBO J 2002 May 15; 21 (10), 2496-506
PMID: 12006501
-
The spindle checkpoint: structural insights into dynamic signalling.
Musacchio A, Hardwick KG
Nat Rev Mol Cell Biol 2002 Oct; 3 (10), 731-41
PMID: 12360190
(click table headers for sorting; Notes column: =Number of Switches, =Number of Interactions)
| Acc., Gene-, Name | Start | End | Subsequence | Logic | #Ev. | Organism | Notes |
|---|---|---|---|---|---|---|---|
| Q12834 CDC20 CDC20_HUMAN |
129 | 137 | DVEEAKILRLSGKPQNAPEG | TP | 3 | Homo sapiens (Human) | |
| Q9Y6D9 MAD1L1 MD1L1_HUMAN |
541 | 549 | DQSRTKVLHMSLNPTSVARQ | TP | 1 | Homo sapiens (Human) | |
| O93289 Fizzy1 O93289_XENLA |
137 | 145 | DMEEAKILRLGGRPQNAPEG | TP | 0 | Xenopus laevis (African clawed frog) | |
| P26309 CDC20 CDC20_YEAST |
201 | 209 | LDMNKRILQYMPEPPKCSSL | TP | 0 | Saccharomyces cerevisiae (Baker"s yeast) | |
| P40957 MAD1 MAD1_YEAST |
581 | 589 | KEKKIRILQLRDGPFIKDQF | TP | 0 | Saccharomyces cerevisiae (Baker"s yeast) | |
| Q24044 fzy Q24044_DROME |
176 | 184 | HNNPLKVVYSIKTPISTKSG | TP | 0 | Drosophila melanogaster (Fruit fly) |
Please cite:
ELM-the Eukaryotic Linear Motif resource-2024 update.
(PMID:37962385)
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement