LIG_EVH1_3
| Accession: | |
|---|---|
| Functional site class: | EVH1 ligands |
| Functional site description: | Proline-rich sequences that bind to the signal transduction modules EVH1. |
| ELMs with same func. site: | LIG_EVH1_1 LIG_EVH1_2 LIG_EVH1_3 |
| ELM Description: | The EVH1_3 motif can be divided into two regions. The N-terminal part comprises two phenylalanines (in several cases substituted by tyrosine and tryptophan, respectively). It is followed by a polyproline region, as for all EVH1-binding motifs, which forms a polyproline helix upon binding. In LIG_EVH1_3 motifs, this region consists of a hydrophobic amino acid, two highly conserved prolines (separated by one position) and an acidic residue (substituted by an arginine in distinct species, eg. yeast). The polyproline region resembles the one of LIG_EVH1_1 motif as it also contains a hydrophobic residue followed by prolines, yet the conserved prolines occupy different positions. The motif is followed by another highly conserved region demonstrating a pattern [RK].YPS. However, it was not included in the ELM regular expression as its significance for EVH1-binding remains unclear. |
| Pattern: | [FY].[FW].....[LMVIF]P.P[DE] |
| Pattern Probability: | 1.855e-07 |
| Present in taxon: | Eukaryota |
| Interaction Domain: |
WH1 (PF00568)
WH1 domain
(Stochiometry: 1 : 1)
|
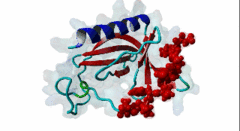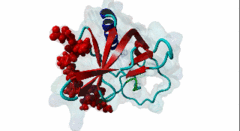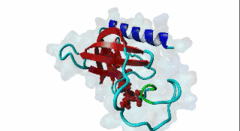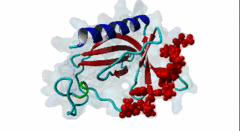
Enabled/vasodilator-stimulated phosphoprotein homology 1 (EVH1) domains are 115 residue protein-protein interaction modules which provide essential links for their host proteins to various signal transduction pathways. Many EVH1-containing proteins are associated closely with actin-based structures and are involved in re-organization of the actin cytoskeleton. EVH1 domains are also present in proteins enriched in neuronal tissue, thus implicating them as potential mediators of synaptic plasticity, linking them to memory formation and learning. EVH1 domains recognize and bind specific proline-rich sequences (PRSs). In general, a small (3-6 residue) core PRS in the target protein binds a 'recognition pocket' on the domain surface. Further affinity- and specificity-increasing interactions are then formed between additional domain epitopes and peptide 'core-flanking' residues. The protein-protein interactions mediated by EVH1 domains are clearly highly important for the regulation of signal transduction events, re-organization of the actin cytoskeleton, and modulation of actin dynamics and actin-based motility. EVH1 domains can be divided into three classes based on the consensus sequences of their target PRS ligands. Single residue peptide substitution experiments have shown the consensus binding motif for class I EVH1 domains to be F-P-X-(hydrophobic)-P. Such sequences are found in a number of mammalian proteins, including the focal adhesion proteins zyxin and vinculin, as well as in the ActA protein of the intracellular pathogen Listeria monocytogenes. Class II EVH1 domains are found in the Homer-Vesl protein family of postsynaptic receptor associated proteins. These recognize a consensus PRS, with a core motif comprising PPxxF, found in the group I mGluRs, IP3Rs, RyRs and the Shank family proteins. The EVH1 domains exclusive for WASP and N-WASP proteins are also called WH1/ WASP-homology 1 domains. The EVH1 class III motifs are found in the verprolin family of proteins. They are known to play a role in WASP/N-WASP- mediated processes (e.g. regulation actin filament dynamics by WIP and WIRE), yet their exact functions remain to be elucidated. The recognition motif of EVH1 class III includes a polyproline region, which is a common feature of all EVH1 motifs, yet in class III it comprises an additional region which contains two highly conserved phenylalanines. |
-
CR16 forms a complex with N-WASP in brain and is a novel member of a
conserved proline-rich actin-binding protein family.
Ho HY, Rohatgi R, Ma L, Kirschner MW
Proc Natl Acad Sci U S A 2001 Sep 25; 98 (20), 11306-11
PMID: 11553796
-
EVH1 domains: structure, function and interactions.
Ball LJ, Jarchau T, Oschkinat H, Walter U
FEBS Lett 2002 Feb 20; 513 (1), 45-52
PMID: 11911879
-
The WASP-binding protein WIRE has a role in the regulation of the actin
filament system downstream of the platelet-derived growth factor receptor.
Aspenstrom P
Exp Cell Res 2002 Sep 10; 279 (1), 21-33
PMID: 12213210
-
The WH1 and EVH1 domains of WASP and Ena/VASP family members bind distinct
sequence motifs.
Zettl M, Way M
Curr Biol 2002 Sep 17; 12 (18), 1617-22
PMID: 12372256
-
Structure of the N-WASP EVH1 domain-WIP complex: insight into the
molecular basis of Wiskott-Aldrich Syndrome.
Volkman BF, Prehoda KE, Scott JA, Peterson FC, Lim WA
Cell 2002 Nov 15; 111 (4), 565-76
PMID: 12437929
-
The mammalian verprolin homologue WIRE participates in receptor-mediated
endocytosis and regulation of the actin filament system by distinct
mechanisms.
Aspenstrom P
Exp Cell Res 2004 Aug 15; 298 (2), 485-98
PMID: 15265696
-
Multiple WASP-interacting protein recognition motifs are required for a
functional interaction with N-WASP.
Peterson FC, Deng Q, Zettl M, Prehoda KE, Lim WA, Way M, Volkman BF
J Biol Chem 2007 Mar 16; 282 (11), 8446-53
PMID: 17229736
(click table headers for sorting; Notes column: =Number of Switches, =Number of Interactions)
| Acc., Gene-, Name | Start | End | Subsequence | Logic | #Ev. | Organism | Notes |
|---|---|---|---|---|---|---|---|
| Q9Z0G8 Wipf3 WIPF3_RAT |
425 | 437 | ESKFTFHSMEDFPPPDEYKP | TP | 4 | Rattus norvegicus (Norway rat) | |
| Q8TF74 WIPF2 WIPF2_HUMAN |
400 | 412 | ESKYSFHPVEDFPAPEEYKH | TP | 8 | Homo sapiens (Human) | |
| O43516 WIPF1 WIPF1_HUMAN |
454 | 466 | ESRFYFHPISDLPPPEPYVQ | TP | 12 | Homo sapiens (Human) |
Please cite:
ELM-the Eukaryotic Linear Motif resource-2024 update.
(PMID:37962385)
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement