| Accession: | |
|---|---|
| Functional site class: | N-degron |
| Functional site description: | The N-end rule pathway regulates protein stability by targeting proteins for ubiquitin-dependent proteasomal degradation. Polyubiquitylation of N-end rule substrates depends on their recognition by N-recognins, specific E3 ubiquitin ligases that use their conserved UBR-box and N-box domains to bind specific N-terminal protein motifs, called N-degrons, in their target proteins. N-degrons are defined by a destabilizing N-terminal residue. Type I destabilizing residues can either occur as primary destabilizing residues, which are positively charged amino acids directly recognized by N-recognins, or as secondary and tertiary destabilizing amino acids, which can be conjugated to a primary destabilizing residue. N-degrons containing type I destabilizing residues are specifically bound by the UBR-box of N-recognins. In contrast, type II destabilizing residues, which comprise bulky hydrophobic amino acids, initiate protein degradation by binding to the N-box of N-recognins. |
| ELMs with same func. site: | DEG_Nend_Nbox_1 DEG_Nend_UBRbox_1 DEG_Nend_UBRbox_2 DEG_Nend_UBRbox_3 DEG_Nend_UBRbox_4 |
| ELM Description: | This class of N-degrons is defined by a type II destabilizing Phe, Tyr, Trp, Leu or Ile residue in the N-terminal position, which is recognized by the N-box of N-recognins. Functional degrons of this class are generated from pre-N-degrons by internal proteolytic cleavage (Varshavsky,2011; Tasaki,2007). Generation by excision of the N-terminal Met on nascent proteins has not yet been investigated, since the known N-terminal Met-aminopeptidases show no activity towards amino acids with larger side chains like Phe, Tyr, Trp, Leu or Ile (Xiao,2010; Varshavsky,2011). It is important to note that the ELM prediction tool will only return internal N-degrons if the sequence of the cleavage product is entered for analysis. Once the N-degron is unmasked, having the Phe, Tyr, Trp, Leu or Ile residue in the N-terminal position, it is recognized by the N-box domain of UBR proteins. The N-box forms a hydrophobic binding pocket that interacts with the N-terminal hydrophobic side chain. In addition, binding involves backbone interactions with the first and second residue of the degron (Tasaki,2012). Bioinformatical analysis and mutational studies showed that the mammalian type II binding site has similarities to ClpS, the ATP-dependent protease adaptor protein of the bacterial proteasomal system, which lacks ubiquitylation (Mogk,2007; Roman-Hernandez,2009) (3G19). The prokaryotic ClpS-mediated proteolysic system was most probably the precursor of the eukaryotic type II N-degron recognition via the N-box (Sriram,2010). The N-box domain is conserved in the UBR1 protein of S. cerevisiae as well as in the mammalian UBR proteins. It is located close to the UBR-box, which specifically binds to type I destabilizing residues. Recent studies suggest an interaction of both domains, since deletion of the UBR-box diminished the binding of the type II substrates. This indicates that the N-box is essential but not sufficient for binding of type II N-degrons (Tasaki,2009). |
| Pattern: | ^M{0,1}[FYLIW][^P] |
| Pattern Probability: | 0.0002302 |
| Present in taxon: | Eukaryota |
| Interaction Domain: |
zf-UBR (PF02207)
Putative zinc finger in N-recognin (UBR box)
(Stochiometry: 1 : 1)
|
The half-life of proteins, which can vary from a few seconds to several days and even years (Varshavsky,2011), is determined by regulated proteolytic degradation. A degradation system common to Eukaryotes is the ubiquitin proteasome system (UPS). Proteins exhibiting degradation signals are recognized and polyubiquitylated by ubiquitin ligases, leading to their subsequent proteasomal degradation. A subset of these degradation signals can be found at the N-terminus of proteins or at the neo-N-terminus of protein cleavage products. These N-terminal motifs, called N-degrons, are recognized by E3 ubiquitin ligases known as N-recognins, which mark their substrates for proteasomal degradation by polyubiquitylation of Lys residues (Bachmair,1986). Studies suggest that this highly conserved N-end rule pathway is the most frequent way of controlled protein degradation, being involved in regulation of G-protein signalling, control of peptide import, regulation of apoptosis, fidelity of chromosome segregation and maintenance of amino acid pools during starvation of cells (Hu,2005). Currently, 12 destabilizing N-terminal residues are known and classified into two groups of N-degrons according to their physicochemical properties. Type I destabilizing residues comprise positively charged Arg and Lys whereas type II residues are amino acids with large hydrophobic side chains including Tyr, Trp, Phe, Leu and Ile (Tasaki,2012). 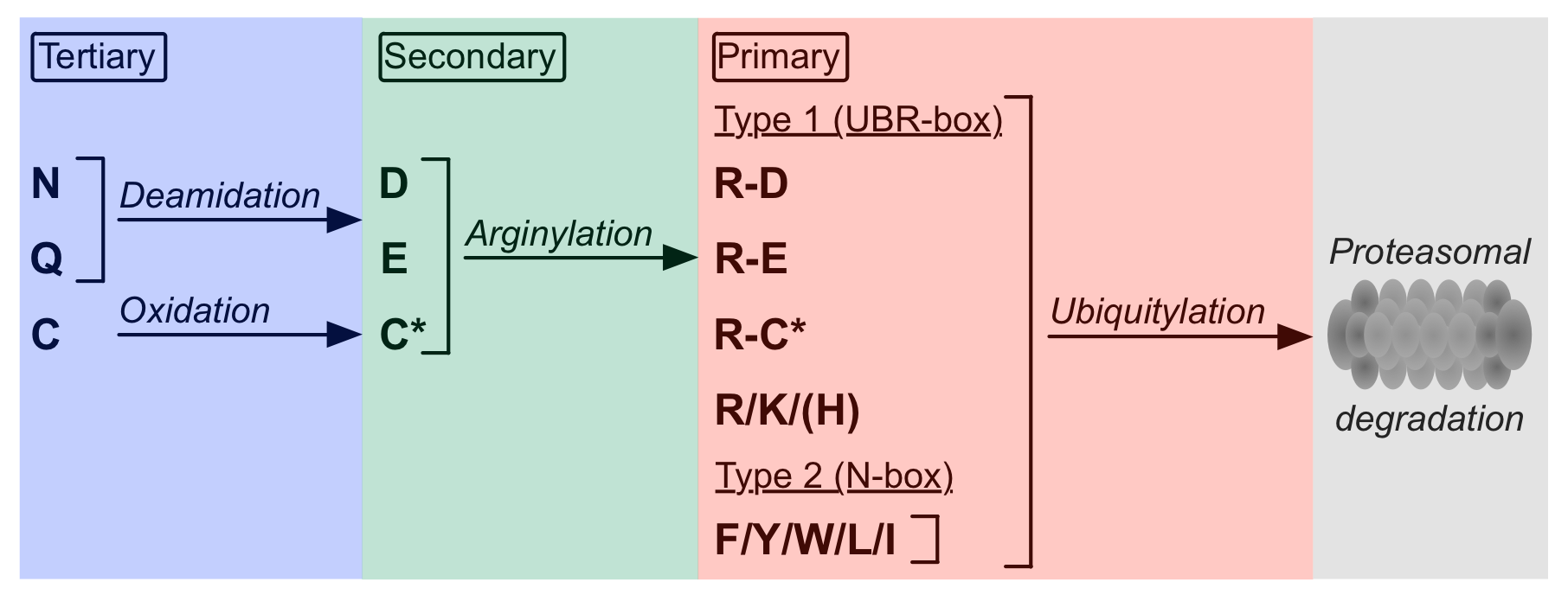 These primary destabilizing residues are directly recognized by N-recognins. In addition, the N-end rule pathway also contains secondary and tertiary destabilizing amino acids, which are destabilizing due to their ability of being conjugated to a primary destabilizing residue. Glu and Asp are secondary destabilizing residues as they can be conjugated to Arg by ATE1-encoded arginyl transferases (R-transferases) (Kwon,1999). These R-transferases are specific for Glu and Asp but cannot arginylate Asn and Gln. Before being arginylated, the tertiary destabilizing residues Asn and Gln are first deamidated by N-terminal amidohydrolases to create the secondary destabilizing residues Asp and Glu. In addition, Cys was also found to function as a tertiary destabilizing residue in higher Eukaryotes but not in Fungi. In the presence of oxygen and nitric oxide, N-terminal Cys gets oxidized to sulfinic (CysO2(H)) or sulfonic (CysO3(H)) acid. As structural mimics of Asp, these can be recognized and arginylated by R-transferases. Due to the requirement of cellular oxygen and nitric oxide, this subset of N-end rule pathway substrates is thought to mediate oxygen sensing in both Mammalia and Viridiplantae systems (Hu,2005; Licausi,2011). These primary destabilizing residues are directly recognized by N-recognins. In addition, the N-end rule pathway also contains secondary and tertiary destabilizing amino acids, which are destabilizing due to their ability of being conjugated to a primary destabilizing residue. Glu and Asp are secondary destabilizing residues as they can be conjugated to Arg by ATE1-encoded arginyl transferases (R-transferases) (Kwon,1999). These R-transferases are specific for Glu and Asp but cannot arginylate Asn and Gln. Before being arginylated, the tertiary destabilizing residues Asn and Gln are first deamidated by N-terminal amidohydrolases to create the secondary destabilizing residues Asp and Glu. In addition, Cys was also found to function as a tertiary destabilizing residue in higher Eukaryotes but not in Fungi. In the presence of oxygen and nitric oxide, N-terminal Cys gets oxidized to sulfinic (CysO2(H)) or sulfonic (CysO3(H)) acid. As structural mimics of Asp, these can be recognized and arginylated by R-transferases. Due to the requirement of cellular oxygen and nitric oxide, this subset of N-end rule pathway substrates is thought to mediate oxygen sensing in both Mammalia and Viridiplantae systems (Hu,2005; Licausi,2011).N-degrons can be generated either by excision of the N-terminal Met or by internal proteolytic cleavage, which is carried out by proteases such as separases and caspases (Varshavsky,2011; Tasaki,2007). Met excision is catalyzed by N-terminal methionine aminopeptidases. Cleavage by these peptidases requires a small residue in the scissile bond C-terminal position. Among the known destabilizing amino acids, only Cys fulfills this requirement, suggesting that most N-degrons are created by internal cleavage (Hu,2005). Once an N-degron is generated, the motif is bound by N-recognins, which initiate the degradation process. N-recognins, also referred to as UBR proteins, are members of the E3 ubiquitin ligase family. The yeast S. cerevisiae expresses one known UBR protein (Ubr1), whereas mammalian genomes encode at least 7 UBR proteins. In plants, PRT1 (Stary,2003) and PRT6 (Garzon,2007) have been identified as N-recognins (Graciet,2010). Canonical UBR proteins contain two motif-binding domains (Tasaki,2009). The UBR-box is composed of two zinc fingers forming a negatively charged binding pocket that specifically interacts with primary type I destabilizing residues (3NY3). The N-box recognizes Type II destabilizing residues via hydrophobic interactions (Tasaki,2012; Choi,2010). Additional domains found in N-recognins are the RING finger domain and the auto-inhibitory domain that catalyze and regulate protein ubiquitylation, respectively. Under physiological conditions UBR proteins build complexes with E2 conjugating enzymes, like the human HR6A and HR6B proteins (Sriram,2011). To date, only few physiological N-end rule substrates have been experimentally characterized, most of which are endoproteolytic protein cleavage products. They can be categorized into 5 different classes according to their destabilizing residue. DEG_Nend_UBRbox_1 contains type I primary destabilizing residues whereas DEG_Nend_UBRbox_2 comprises the secondary destabilizing residues Glu and Asp. N-degrons depending on tertiary destabilizing residues are described in DEG_Nend_UBRbox_3 (Asn or Gln) and DEG_Nend_UBRbox_4 (Cys), the latter of which is not functional in Fungi. N-degrons containing type II destabilizing residues are captured in DEG_Nend_Nbox_1. |
Please cite:
ELM-the Eukaryotic Linear Motif resource-2024 update.
(PMID:37962385)
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
