| Accession | Acc. Gene-, Name | Start | End | Subsequence | Logic | PDB | Organism | Length |
|---|---|---|---|---|---|---|---|---|
| ELMI001705 | O43516 WIPF1 WIPF1_HUMAN |
454 | 466 | ESRFYFHPISDLPPPEPYVQ | TP |
1MKE 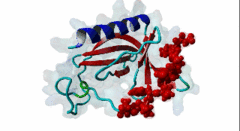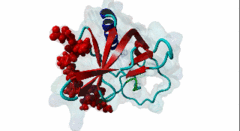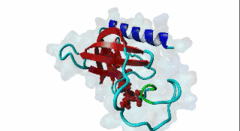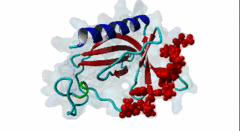
|
Homo sapiens (Human) | 503 |
![]() Instance evidence
Instance evidence
| Evidence class | PSI-MI | Method | BioSource | PubMed | Logic | Reliability | Notes |
|---|---|---|---|---|---|---|---|
| experimental | MI:0113 | western blot | in vitro | Volkman,2002 | support | certain | ParticipantIdentification FeatureDetection |
| experimental | MI:0077 | nuclear magnetic resonance | in vitro | Volkman,2002 | support | certain | InteractionDetection FeatureDetection |
| experimental | MI:0074 | mutation analysis | in vivo/in vitro | Volkman,2002 | support | certain | FeatureDetection |
| experimental | MI:0105 | structure based prediction | in silico | Volkman,2002 | support | certain | InteractionDetection |
| experimental | MI:0074 | mutation analysis | in vivo/in vitro | Peterson,2007 | support | certain | FeatureDetection |
| experimental | MI:0077 | nuclear magnetic resonance | in vitro | Peterson,2007 | support | certain | InteractionDetection FeatureDetection |
| experimental | MI:0105 | structure based prediction | in silico | Peterson,2007 | support | certain | InteractionDetection |
| experimental | MI:0005 | alanine scanning | in vivo/in vitro/in silico | Peterson,2007 | support | certain | FeatureDetection |
| experimental | MI:0367 | green fluorescent protein tag | in vivo | Zettl,2002 | support | certain | |
| experimental | MI:0005 | alanine scanning | in vivo/in vitro/in silico | Zettl,2002 | support | certain | FeatureDetection |
| experimental | MI:0074 | mutation analysis | in vivo/in vitro | Zettl,2002 | support | certain | FeatureDetection |
| experimental | MI:0096 | pull down | in vivo/in vitro | Zettl,2002 | support | certain | InteractionDetection |
![]() Interactions
Interactions
| Uniprot Id | Domain family | Domain Start | Domain End | Affinity Min/Max (µMol) | Notes |
|---|---|---|---|---|---|
| (O08816) WASL_RAT | PF00568
(WH1)
WH1 domain |
28 | 135 | [mitab][xml] |
![]() Pathways
Pathways
Please cite:
ELM-the Eukaryotic Linear Motif resource-2024 update.
(PMID:37962385)
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
