| Accession: | |
|---|---|
| Functional site class: | MYND domain binding motif. |
| Functional site description: | The MYND domain is a zinc binding domain that is involved in protein-protein interactions mainly in the context of transcriptional regulation. It is named after Myeloid, Nervy, and DEAF-1, which are the three most characterized proteins that contain the MYND domain. Only a small number of MYND domain containing proteins have been identified and they are involved in various biological processes such as cell proliferation, apoptosis, adhesion, migration, and tumorigenesis and oxygen homeostasis. MYND domain typically binds a proline-rich motif in their interacting partners, however they have different binding specificities. |
| ELMs with same func. site: | LIG_MYND_1 LIG_MYND_2 LIG_MYND_3 |
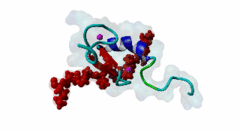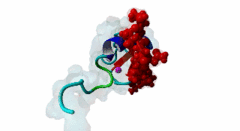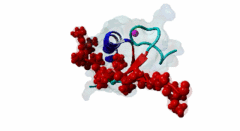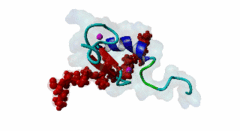
| ELM Description: | The MYND domain (PF01753) of AML1/ETO (Q06455) recognizes co-repressor proteins through a PPxLI motif. Solution structure of SMRT (Q9Y618) peptide bound to MYND domain has revealed that SMRT/N-Cor peptide (1113-PPPLI-1117) forms an anti-parallel beta-sheet against a hydrophobic pocket of the MYND domain (2ODD) (Liu,2007). The Leu1116, Ile1117, and Ser1118 residues of SMRT are involved in this beta-sheet. Furthermore, Pro1113 of SMRT and Trp692 of MYND are packed together, while Pro1114 and Leu1116 residues of SMRT form hydrogen bonds with Ser675 and Gln688 of MYND. Mutations of Trp692, Gln688, or Ser675 to alanine have been shown to disrupt binding of MYND to SMRT, while the structure of MYND was preserved. This ‘PPPLI’ peptide is conserved in NCor1 (O75376) (1033-PPPLI-1037) and mediates the same interaction with MYND domain. Another similar peptide, SMRT (1674-PPYLI-1678), also binds to MYND domain, however with a weaker affinity. Thus, the core motif is constructred as ‘PPxLI’ instead of the commonly mentioned version ‘PPPLI’, because the second peptide of SMRT has a Tyr instead of Pro at the third position and in the solution structure of the first peptide of SMRT, no role has been attributed to this position for binding to the MYND domain. In the first peptide of SMRT, the solution structure reveals a certain role for Ser1118, which is part of the anti-parallel beta sheet. This Ser is well conserved in SMRT orthologs, however, NCor1 has a conserved Arg instead of Ser at this position. Thus, this position was not included in the core motif definition. |
| Pattern: | PP.LI |
| Pattern Probability: | 0.0000175 |
| Present in taxon: | Metazoa |
| Interaction Domain: |
zf-MYND (PF01753)
MYND finger
(Stochiometry: 1 : 1)
|
The MYND domain is a cysteine-rich structure present in proteins generally implicated in gene regulation and associated with cancers. Some of the MYND domain containing proteins are BS69, Nervy (a transcriptional corepressor), the chimeric fusion protein of acute myelogenous leukemia (AML) and ETO (a nuclear protein that interacts with corepressor molecules), the bone morphogenesis protein receptor-associated molecule1 (BRAM1), SET and MYND domain-containing proteins (SMYD), RACK7, PDCD2, deformed epidermal auto regulatory factor-1 (DEAF-1). The structures of the MYND domain in these proteins demonstrate a tandem zinc-binding motif organized in cross-brace topology (Kateb,2013). A variant of MYND domain is also found to exist in prolyl hydroxylase domain protein 2 (PHD2) that catalyzes proline hydroxylation of HIFα subunits. Prolyl hydroxylation is an ancient mechanism to transduce changes in oxygen concentration to changes in cellular function. Among the three PHD isoforms, PHD2 is a distinct one as it is predicted to have a MYND-type zinc finger at its N-terminal. It is also the one that is most closely related to the single ancestral PHD and has the capacity to hydroxylate HIF-1α, which is connected to HSP90 pathway (Huang,2002). Depending upon the variations in amino acid sequence of MYND domain, there exist different binding specificities for MYND domain containing proteins and it acts as a scaffold for different cellular and viral protein-protein interactions. MYND domains usually recognize proline-rich motifs in their partners. The MYND domain of BS69 has been shown to bind viral tumour antigens E1A and EBNA2 as well as with cellular partners such as MGA through a PxLxP motif (Ansieau,2002). The MYND domain of BS69 contains a set of positively charged residues at its C-terminus, which is crucial for interaction with PxLxP ligands. Other MYND domain containing proteins such as RACK7, ETO and DEAF-1, which lack these charged residues fail to interact with the PxLxP motif. The co-repressor-binding MYND domain-containing proteins such as DEAF-1 and ETO have higher degree of sequence similarity and similar binding mode. They can bind to another proline-rich motif PPPLI (Liu,2007). The MYND-type Zinc Finger of PHD2 binds a PxLE motif, which is so far found to exist in HSP90 co-chaperones p23 and FKBP38 (Song,2013). |
(click table headers for sorting; Notes column: =Number of Switches, =Number of Interactions)
| Acc., Gene-, Name | Start | End | Subsequence | Logic | #Ev. | Organism | Notes |
|---|---|---|---|---|---|---|---|
| Q9Y618 NCOR2 NCOR2_HUMAN |
1113 | 1117 | RPPTISNPPPLISSAKHPSV | TP | 4 | Homo sapiens (Human) | |
| O75376 NCOR1 NCOR1_HUMAN |
1033 | 1037 | TTRPTRPPPPLIPSSKTTVA | TP | 3 | Homo sapiens (Human) | |
| Q9Y618 NCOR2 NCOR2_HUMAN |
1674 | 1678 | PTYPHLYPPYLIRGYPDTAA | TP | 2 | Homo sapiens (Human) |
Please cite:
ELM-the Eukaryotic Linear Motif resource-2024 update.
(PMID:37962385)
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement
ELM data can be downloaded & distributed for non-commercial use according to the ELM Software License Agreement